Publications
. A cyclical fast iterative method for simulating reentries in cardiac electrophysiology using an eikonal-based model. Engineering with Computers, 296–314, January 2025. URL https://doi.org/10.1007/s00366-024-02094-9. Online first.
. Differential effects of mechano-electric feedback mechanisms on whole-heart activation, repolarization, and tension. J. Physiol., 20pp., January 2024. URL https://doi.org/10.1113/JP285022. Online first. [files]
. Numerical evaluation of elasto-mechanical and visco-elastic electro-mechanical models of the human heart. GAMM-Mitt., 46(3-4):e202370010, September 2023. URL https://doi.org/10.1002/gamm.202370010.
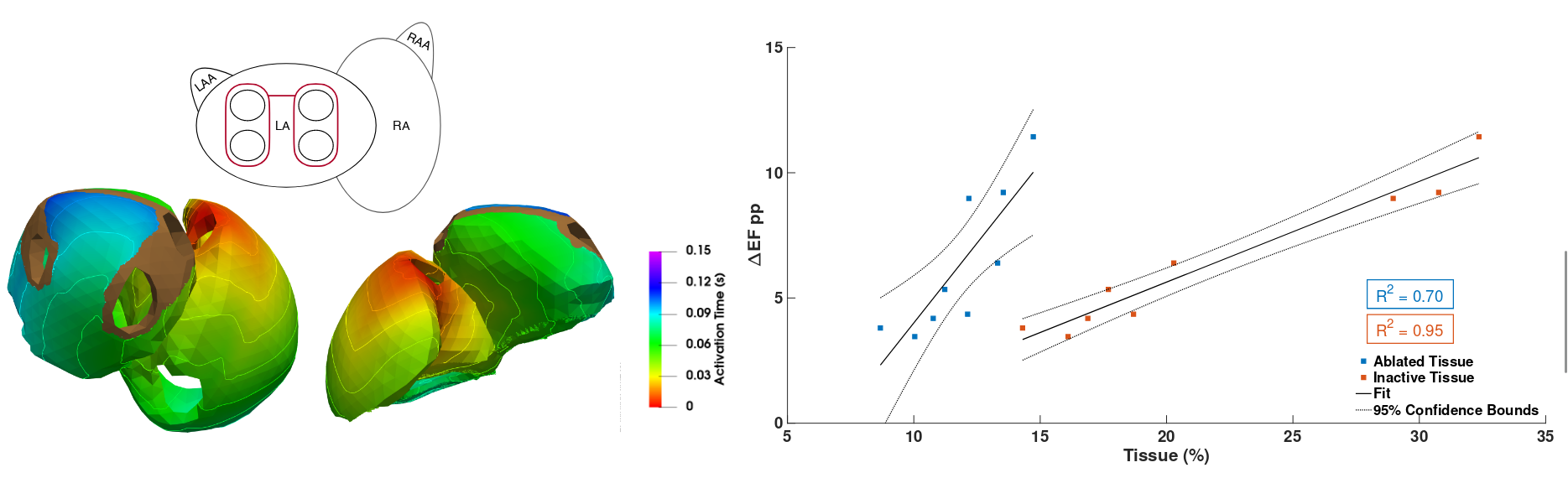
. The impact of standard ablation strategies for atrial fibrillation on cardiovascular performance in a four-chamber heart model. Cardiovasc. Eng. Technol., 14(2):296–314, April 2023. URL https://doi.org/10.1007/s13239-022-00651-1. [preprint]
. Efficient time splitting schemes for the monodomain equation in cardiac electrophysiology. Int. J. Numer. Method. Biomed. Eng., 39(2):e3666, February 2023. URL https://doi.org/10.1002/cnm.3666. [preprint]
. The reverse mode of the Na$^+$/Ca$^{2+}$ exchanger contributes to the pacemaker mechanism in rabbit sinus node cells. Sci. Rep., 12(1):21830, December 2022. URL https://doi.org/10.1038/s41598-022-25574-8.
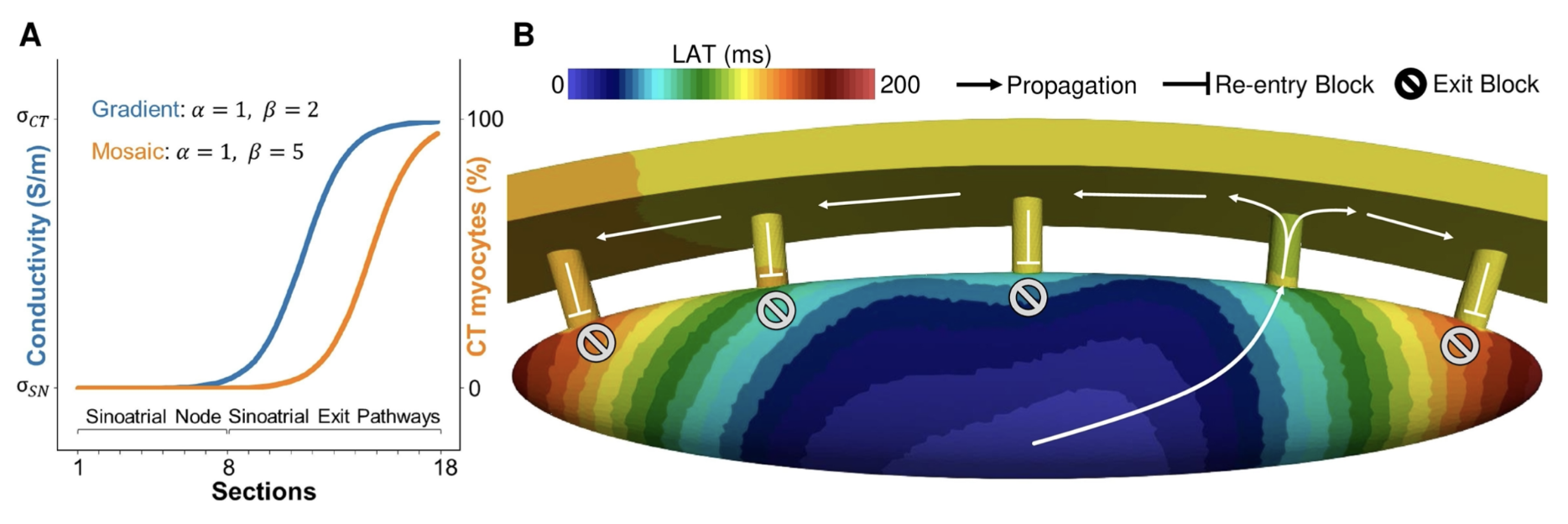
. Characterization of the pace-and-drive capacity of the human sinoatrial node: a 3D in silico study. Biophys. J., 121(22):4247–4259, November 2022. URL https://doi.org/10.1016/j.bpj.2022.10.020. [files]
. Dyssynchronous left ventricular activation is insufficient for the breakdown of wringing rotation. Front. Physiol., 13:838038, May 2022. URL https://doi.org/10.3389/fphar.2022.838038.
. Causes of altered ventricular mechanics in hypertrophic cardiomyopathy: an in-silico study. BioMed. Eng. OnLine, 20:69, July 2021. URL https://doi.org/10.1186/s12938-021-00900-9.
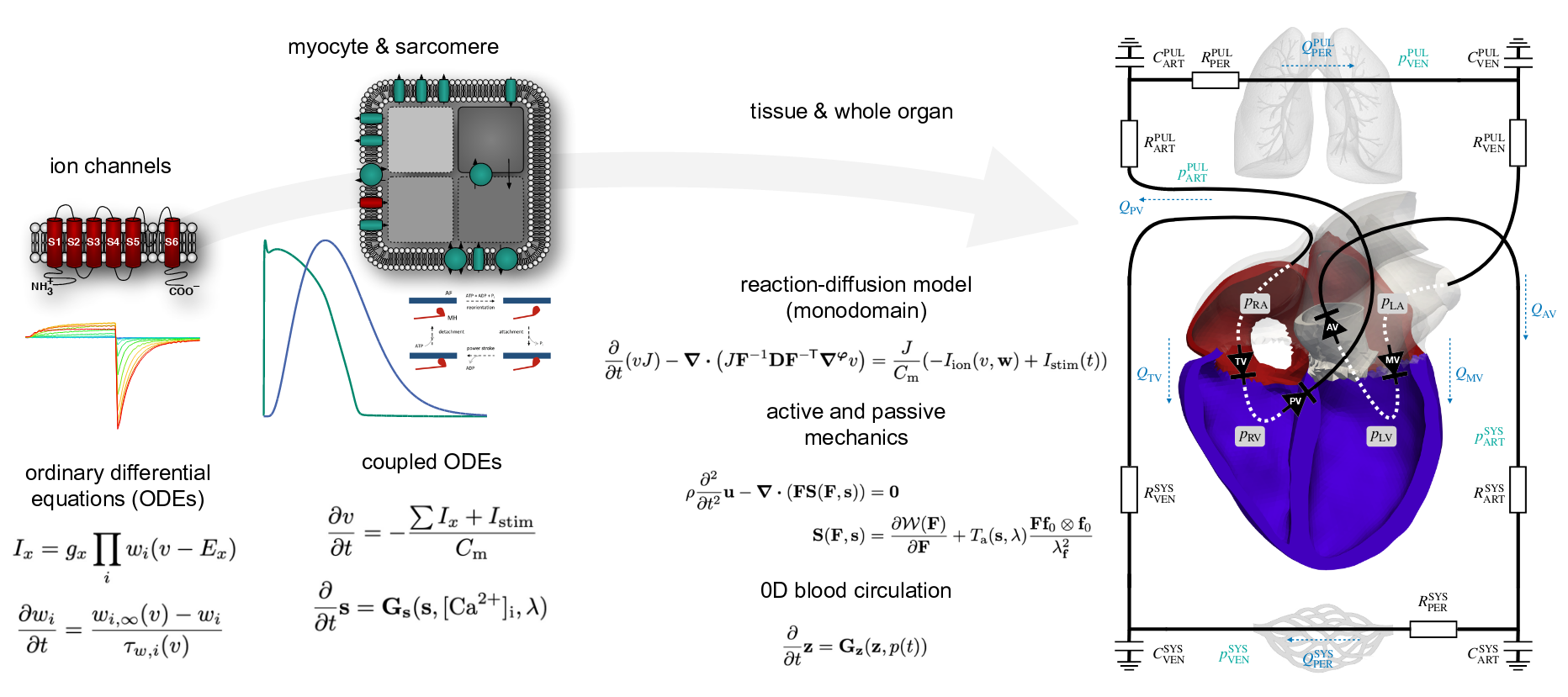
. Electro-mechanical whole-heart digital twins: A fully coupled multi-physics approach. Mathematics, 9(11):1247, May 2021. URL https://doi.org/10.3390/math9111247.
. Cycle length statistics during human atrial fibrillation reveal refractory properties of the underlying substrate – a combined in silico and clinical test of concept study. Europace, 23(Supplement_1):i133–i142, March 2021. URL https://doi.org/10.1093/europace/euaa404.
. Consequences of using an orthotropic stress tensor for left ventricular systole. In 2020 Computing in Cardiology (CinC), pages 1–4, Rimini, Italy, September 2020.
. The cardiac pacemaker story — Fundamental role of the Na${}^{+}$/Ca${}^{2+}$ exchanger in spontaneous automaticiy. Front. Pharmacol., 11:516, April 2020. URL https://doi.org/10.3389/fphar.2020.00516.
. Mapping and removing the ventricular far field component in unipolar atrial electrograms. IEEE Trans. Biomed. Eng., 67(10):2905–2915, February 2020. URL https://doi.org/10.1109/TBME.2020.2973471.
. Novel Na${}^{+}$/Ca${}^{2+}$ exchanger inhibitor ORM-10962 supports coupled function of funny-current and Na${}^{+}$/Ca${}^{2+}$ exchanger in pacemaking of rabbit sinus node tissue. Front. Pharmacol., 10:1632, January 2020. URL https://doi.org/10.3389/fphar.2019.01632.
. Hypocalcemia-induced slowing of human sinus node pacemaking. Biophys. J., 117(12):2244–2254, December 2019. URL https://doi.org/10.1016/j.bpj.2019.07.037.
. Inter-species differences in the response of sinus node cellular pacemaking to changes of extracellular calcium. In 41st Annual International Conference, pages 1875–1878, October 2019. IEEE Engineering in Medicine and Biology Society (EMBC), IEEE.
. Observation guided systematic reduction of a detailed human ventricular cell model. In 2019 Computing in Cardiology (CinC), volume 46, pages 1–4, February 2019.
. Patient-specific identification of atrial flutter vulnerability—A computational approach to reveal latent reentry pathways. Front. Physiol., 9:1910, January 2019. URL https://doi.org/10.3389/fphys.2018.01910.
. Influence of left atrial size on P-wave morphology: differential effects of dilation and hypertrophy. Europace, 20(suppl_3):iii36–iii44, November 2018. URL https://doi.org/10.1093/europace/euy231.
. Automatic identification of reentry mechanisms and critical sites during atrial tachycardia by analyzing areas of activity. IEEE Trans. Biomed. Eng., 65(10):2334–2344, October 2018. URL https://doi.org/10.1109/TBME.2018.2794321.
. Electro-mechanical delay in the human heart: A study on a simple geometry. In Computing in Cardiology, volume 45, pages 1–4, September 2018. IEEE.
. A computational framework to benchmark basket catheter guided ablation in atrial fibrillation. Front. Physiol., 9:1251, September 2018. URL https://doi.org/10.3389/fphys.2018.01251.
. Novel electrocardiographic criteria for real-time assessment of anterior mitral line block: "`V1 Jump"' and "`V1 Delay"'. J. Am. Coll. Cardiol. EP, 4(7):920–932, July 2018. URL https://doi.org/10.1016/j.jacep.2018.03.007.
. Commentary: Virtual in-silico modeling gided catheter ablation predicts effective linear ablation lesion set for longstanding persistent atrial fibrillation: Multicenter prospective randomized study. Front. Physiol., 8:1113, December 2017. URL https://doi.org/10.3389/fphys.2017.01113.
. Parameter estimation of ion current formulations requires hybrid optimization approach to be both accurate and reliable. Front. Bioeng. Biotechnol., 3:209, January 2016. URL https://doi.org/10.3389/fbioe.2015.00209.
Theses
. Das Monodomain-Modell für die Erregungsausbreitung im menschlichen Herz und seine effiziente numerische Approximation. PhD thesis, Karlsruhe Institute of Technology (KIT), October 2024.
. Personalized electromechanical modeling of the human heart: challenges and opportunities for the simulation of pathophysiological scenarios. PhD thesis, Karlsruhe Institute of Technology (KIT), May 2022.
. In silico modeling, simulation and optimization of human cardiac motion. PhD thesis, Karlsruhe Institute of Technology (KIT), February 2022.
. Simulation of cardiac electrophysiology and biomechanics: from model development to clinical translation. Habilitation thesis, Karlsruhe Institute of Technology (KIT), December 2021.
. Model based estimation of the elastomechanical properties of the human heart. PhD thesis, Karlsruhe Institute of Technology (KIT), July 2021.
. A multiscale in silico study to characterize the atrial electrical activity of patients with atrial fibrillation: a translational study to guide ablation therapy. PhD thesis, Karlsruhe Institute of Technology (KIT), June 2021.
Other references
. Integrated heart—coupling multiscale and multiphysics models for the simulation of the cardiac function. Comput. Methods Appl. Mech. Engrg., 314:345–407, February 2017. URL https://doi.org/10.1016/j.cma.2016.05.031.
. Influence of the earliest right atrial activation site and its proximity to interatrial connections on P-wave morphology. Europace, 18(suppl_4):iv35–iv43, December 2016. URL https://doi.org/10.1093/europace/euw349.
. Operator splitting. In R. Glowinski, S. J. Osher, and W. Yin, editors, Splitting methods in communication, imaging, science, and engineering, pages 95–114. Springer, Cham, December 2016.
. Verification of cardiac mechanics software: benchmark problems and solutions for testing active and passive material behaviour. Proc. R. Soc. A, 471(2184):20150641, December 2015. URL https://doi.org/10.1098/rspa.2015.0641.
. Sensitivity study of fiber orientation on stroke volume in the human left ventricle. In Computing in Cardiology, volume 41, pages 1–4, September 2014. IEEE.
. Simulation of the contraction of the ventricles in a human heart model including atria and pericardium. Biomech. Model. Mechanobiol., 13:627–641, June 2014. URL https://doi.org/10.1007/s10237-013-0523-y.
. Arrhythmic potency of human ether-à-go-go-related gene mutations L532P and N588K in a computational model of human atrial myocytes. Europace, 16(3):435–443, March 2014. URL https://doi.org/10.1093/europace/eut375.
. Alternans and spiral breakup in a human ventricular tissue model. Am. J. Physiol. Heart. Circ. Physiol., 291(3):H1088–H1100, September 2006. URL https://doi.org/10.1152/ajpheart.00109.2006.